Abstract
A comparison of exact and approximate nearest neighbor search methods, with emphasis on the algorithmic and mathematical foundations of Hierarchical Navigable Small World graphs (HNSW) and the Inverted File Index with Product Quantization (IVF-PQ). This post formalizes the approximate nearest neighbor problem, derives complexity bounds, analyzes memory footprints, and provides worked numerical examples on datasets of practical scale. The goal is to make precise why these methods achieve sub-linear query times and orders-of-magnitude memory reductions relative to brute-force search.
1. The Nearest Neighbor Problem
Let be a dataset of vectors in , and let be a distance function (typically Euclidean or inner product). Given a query vector and an integer , the exact \(k\)-nearest neighbor problem asks for the set
that is, the vectors in closest to under .
The approximate variant relaxes this: return a set such that, with high probability, for every and corresponding true neighbor ,
for some approximation factor . In practice the community measures quality via recall@k, the fraction of true top- neighbors returned by the approximate algorithm.
2. Brute Force Baseline
The naive approach computes for every and selects the smallest distances.
Time complexity. Each distance computation in costs , so the total is
Memory. Storing the full dataset in float32 requires
Worked example. For vectors of dimension :
- Distance computations per query: FLOPs.
- Memory: bytes GB.
At a throughput of roughly FLOPs/s on a modern CPU core, this yields latencies on the order of 77 ms per query — tolerable for small workloads, but untenable at scale. The two families of methods described next reduce either the number of distance computations, the cost per computation, or both.
3. HNSW (Hierarchical Navigable Small World)
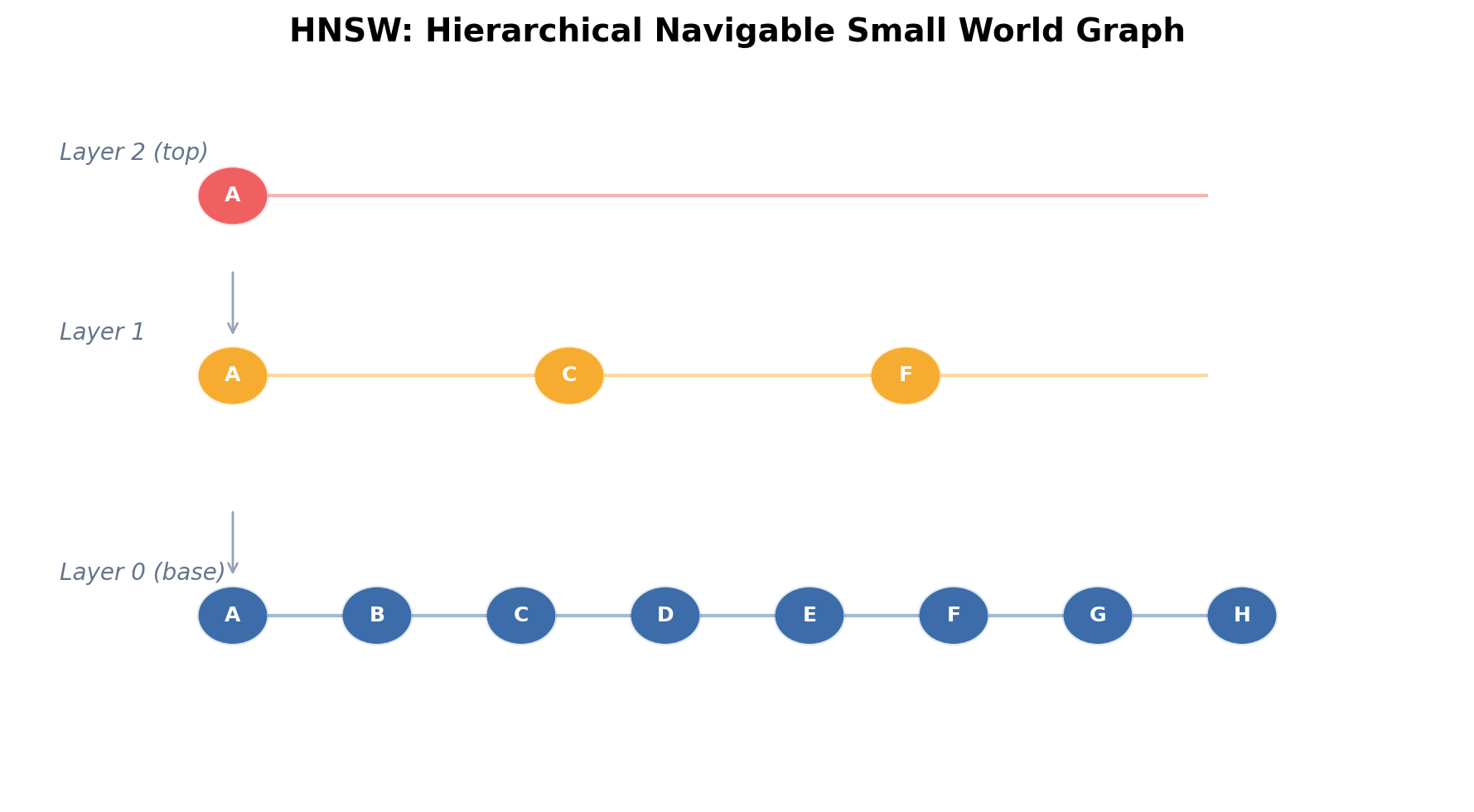
3.1 Navigable Small-World Graphs
A navigable small-world (NSW) graph places every vector as a node and connects each to a set of neighbors such that greedy routing converges quickly. The key structural property is that the graph possesses both short-range links (connecting nearby vectors) and long-range links (enabling logarithmic-hop traversal).
HNSW, introduced by Malkov and Yashunin (2018), layers multiple NSW graphs in a hierarchy. Layers are numbered . Layer 0 contains all nodes; higher layers contain exponentially fewer nodes.
3.2 Layer Assignment
When inserting element , its maximum layer is drawn as
where is a normalization constant, typically set to , with being the maximum number of connections per node. The probability that a node appears at layer or above is
Setting yields an expected number of nodes at layer of
so the expected number of layers is
3.3 Greedy Search
Search begins at the top layer from a fixed entry point. At each layer, a greedy procedure expands a beam (priority queue) of width — called efSearch — by iterating over each candidate's neighbors and keeping the closest vectors found so far. When no improvement is possible at layer , the current best candidate set descends to layer . At layer 0, the procedure terminates and returns the top vectors from the beam.
3.4 Complexity Analysis
At each layer, greedy search visits nodes (expanding candidates, each with neighbors). Across layers, the total number of distance computations is
For fixed and , this is — exponentially better than brute force.
3.5 Recall–Latency Tradeoff
The parameter directly governs the recall–latency tradeoff. Larger means more candidates are evaluated, increasing recall at the cost of more distance computations. Empirically, on 1M-scale datasets with and :
| Recall@10 | Relative latency | |
|---|---|---|
| 16 | ~0.85 | 1× |
| 64 | ~0.95 | 3.5× |
| 256 | ~0.99 | 12× |
Even at , total distance computations are on the order of , a factor of roughly 50× fewer than brute force.
3.6 Memory
HNSW stores the full-precision vectors plus the graph. Graph overhead per node is approximately bytes (storing neighbor IDs as int32). With and layers per node on average, this is roughly 83 bytes per vector — modest compared to the vector storage itself. Total memory for the worked example:
HNSW reduces *compute* dramatically but provides little *memory* savings over brute force. This motivates quantization-based approaches.
4. IVF (Inverted File Index)
4.1 Voronoi Partitioning
The Inverted File Index partitions the dataset into clusters via -means (or a similar algorithm). Let be the centroids. Each vector is assigned to its nearest centroid:
This induces a Voronoi partition of . Each cell stores an inverted list of its members.
4.2 Query Procedure
At query time, the algorithm first computes for all and selects the closest centroids. It then performs exhaustive search only within those inverted lists:
The first term is the centroid comparison cost; the second is the exhaustive scan within selected cells. When , the second term becomes .
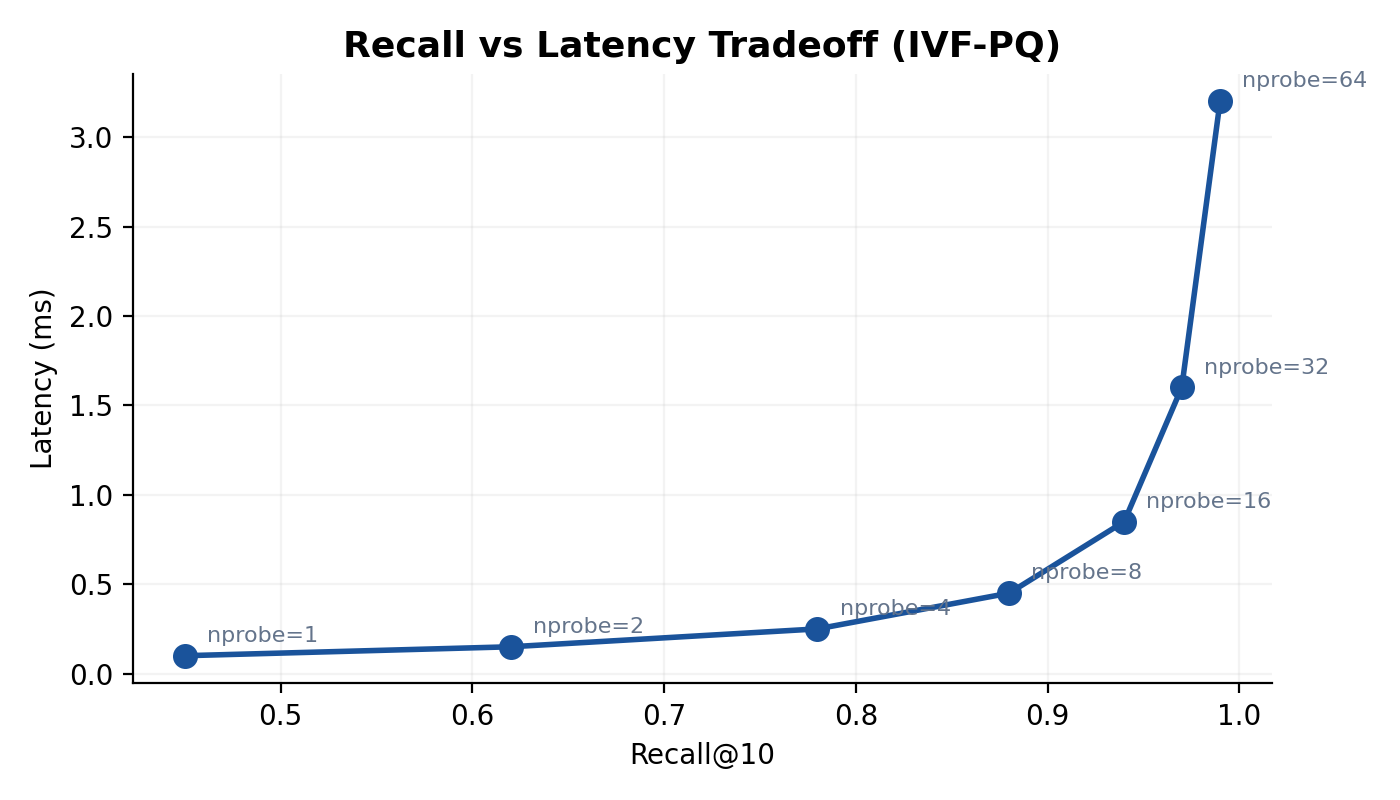
4.3 Recall as a Function of
Under the simplifying assumption that true nearest neighbors are uniformly distributed across Voronoi cells, the probability that a given true neighbor falls within the probed cells is approximately
For independent neighbors, the expected recall is
In practice, neighbors cluster near cell boundaries, so the true recall is higher than this lower bound for reasonable values of . Typical configurations use to and to , achieving recall@10 above 0.90 while scanning fewer than 5% of the dataset.
4.4 Worked Example
With , , , :
- Vectors per cell: .
- Vectors scanned: .
- Distance computations: .
- Speedup vs. brute force: .
Memory remains since the raw vectors are still stored.
5. Product Quantization
5.1 Subspace Decomposition
Product Quantization (PQ), introduced by Jégou et al. (2011), compresses each -dimensional vector into a short code. The vector space is decomposed into disjoint subspaces of dimension each:
5.2 Codebook Learning
For each subspace , a codebook of centroids is learned via -means on the projected sub-vectors . Each sub-vector is then replaced by its nearest centroid index:
The PQ code for vector is the tuple of indices .
5.3 Memory Per Vector
Each index requires bits. With the standard choice , each index is exactly 1 byte. The compressed representation of a single vector requires
5.4 Worked Example: Memory Savings
For , , (subspace dimension = 8), :
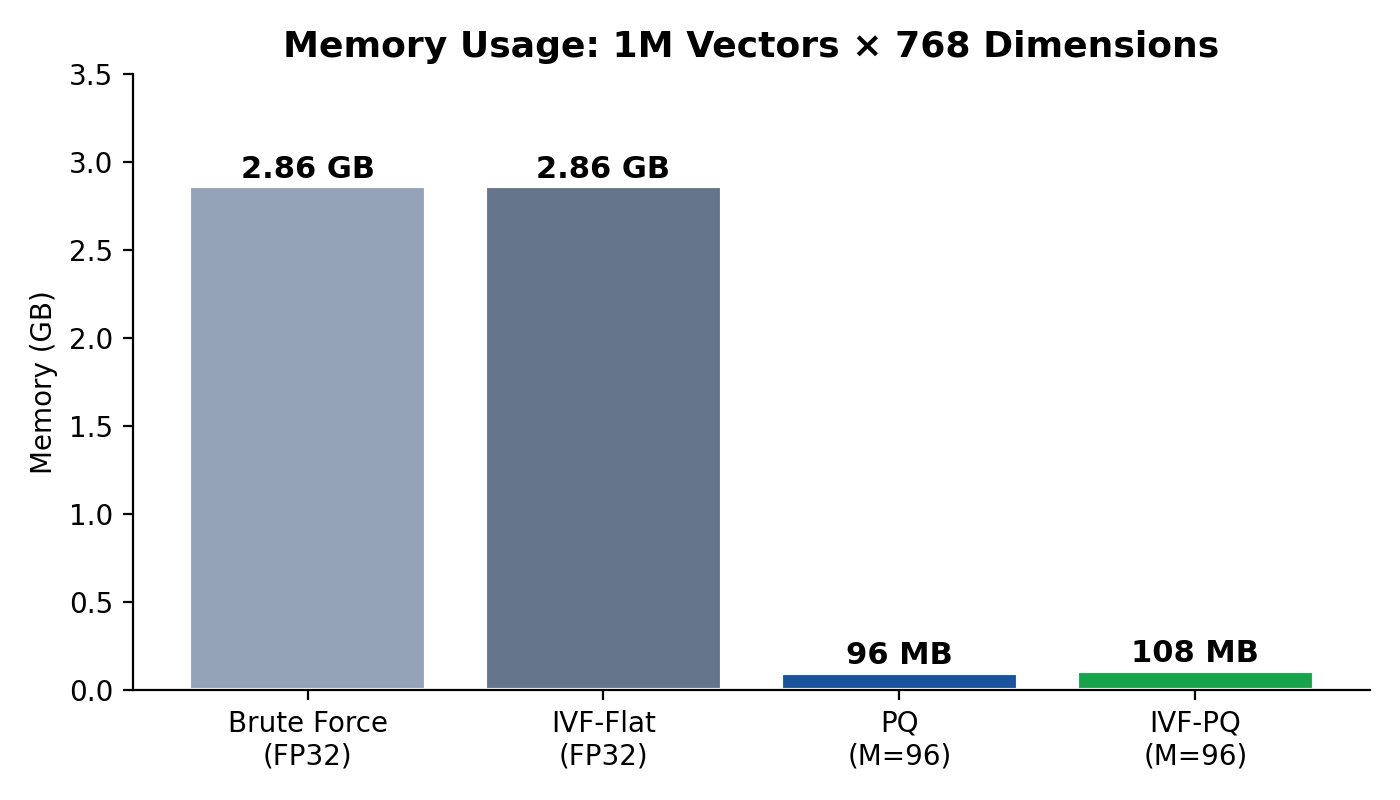
- Raw storage: GB.
- PQ storage: MB.
- Compression ratio: 30.7×.
5.5 Asymmetric Distance Computation (ADC)
At query time, exact distance from query to a database vector (represented by its PQ code) is approximated via asymmetric distance computation. The key insight: precompute a lookup table of sub-vector distances.
For each subspace and each centroid , compute
This produces entries — a table of shape . The approximate squared distance from to is then
Each distance requires only table lookups and additions — no floating-point multiplications.
Lookup table construction cost: FLOPs — independent of .
Per-vector distance cost: lookups + additions .
For the worked example ():
- Table construction: FLOPs (one-time per query).
- Per-vector distance: 96 lookups.
- Total for scanning all 1M vectors: operations.
- Compare to brute force: FLOPs — an 8× reduction in compute, on top of the 30× memory reduction.
6. IVF-PQ Combined
6.1 Two-Stage Pipeline
IVF-PQ combines both ideas into a two-stage pipeline:
- Coarse quantization (IVF): Partition the dataset into Voronoi cells. At query time, select the nearest cells.
- Fine quantization (PQ): Within the selected cells, vectors are stored as PQ codes. Distances are computed via ADC.
An important refinement: rather than quantizing the raw vectors, IVF-PQ typically stores and quantizes the residuals
where is the coarse centroid. Since residuals have smaller magnitude and lower variance than the original vectors, PQ achieves higher fidelity on them.
6.2 Query Procedure
Given query :
- Compute for all centroids. Select top .
- For each selected centroid , compute the query residual .
- Build an ADC lookup table for .
- Scan all PQ codes in cell , computing approximate distances via ADC.
- Merge results across probed cells, return top .
6.3 Compute Analysis
Let be the total number of vectors scanned. The computational cost breaks down as:
6.4 Memory Analysis
The index stores:
- Centroids: bytes.
- PQ codebooks: bytes.
- PQ codes: bytes.
- Vector IDs: bytes (int64).
Total:
6.5 Worked Example: Full Pipeline
Parameters: , , , , , .
Memory:
| Component | Size |
|---|---|
| PQ codes () | MB |
| Vector IDs () | 8 MB |
| Centroids () | 3.0 MB |
| PQ codebooks () | 0.75 MB |
| Total | ~108 MB |
Compared to brute-force storage of 2.86 GB, this represents a 26.5× compression.
Compute per query:
| Stage | Operations |
|---|---|
| Coarse search | |
| Table construction | |
| ADC scan ( vectors lookups) | |
| Total | ~5.4 × 10\(^6\) |
Compared to brute-force (), the speedup is approximately 143×, while the memory footprint has been reduced by a factor of 26.5.
7. Practical Tradeoffs
Accuracy vs. Speed
Both HNSW and IVF-PQ trade exactness for speed, but they occupy different points in the design space:
- HNSW achieves very high recall () with moderate compute overhead. Its graph structure makes it well-suited for latency-critical applications where memory is abundant.
- IVF-PQ sacrifices some recall (typically 0.85–0.95 depending on parameters) in exchange for dramatic memory savings. It is the method of choice when the dataset does not fit in RAM at full precision.
Memory vs. Recall
The relationship between PQ parameters and recall is governed by the quantization distortion. The expected squared distortion per subspace is bounded by the -means quantization error:
where is the optimal quantization error for a -dimensional distribution with centroids. Increasing (more subspaces, each of lower dimension) or increasing reduces distortion but increases code size. In practice, the subspace dimension between 4 and 16 offers the best balance.
Hybrid Approaches
Modern systems often combine both methods. A common architecture uses IVF-PQ as a first-pass retrieval stage, followed by re-ranking with exact distances on the original (or HNSW-indexed) vectors. This two-stage pattern extracts the memory efficiency of PQ and the precision of full-vector comparison:
- IVF-PQ retrieves the top candidates (e.g., ).
- Original vectors for these candidates are fetched and re-ranked.
- The final top are returned.
The re-ranking step costs , which is negligible for moderate .
Scaling Considerations
| Metric | Brute Force | HNSW | IVF-PQ |
|---|---|---|---|
| Query time | |||
| Memory | |||
| Index build | * | ||
| Update support | Trivial | Incremental | Requires retraining |
\* denotes -means iterations.
HNSW supports incremental insertion, making it suitable for streaming workloads. IVF-PQ requires training on a representative sample and is better suited for static or batch-updated corpora.
8. Conclusion
The transition from brute-force to approximate nearest neighbor search is not a single trick but a composition of orthogonal ideas. HNSW reduces the *number* of distance computations from to by organizing vectors in a multi-layer navigable graph. IVF reduces the search scope to a fraction of the dataset via Voronoi partitioning. Product Quantization reduces the *cost* of each distance computation and the *memory* per vector by compressing -dimensional vectors into -byte codes. IVF-PQ combines these last two into a system that can index a billion vectors in tens of gigabytes of RAM with query latencies in the low milliseconds.
Understanding the mathematical underpinnings — layer probability distributions, Voronoi cell geometry, subspace quantization distortion, and asymmetric distance computation — is essential for tuning these systems in production. The parameters , , , , and the PQ subspace configuration are not arbitrary knobs; they are direct consequences of the tradeoffs formalized above.
References
- Malkov, Y. A., & Yashunin, D. A. (2018). Efficient and Robust Approximate Nearest Neighbor Search Using Hierarchical Navigable Small World Graphs. *IEEE Transactions on Pattern Analysis and Machine Intelligence*, 42(4), 824–836.
- Jégou, H., Douze, M., & Schmid, C. (2011). Product Quantization for Nearest Neighbor Search. *IEEE Transactions on Pattern Analysis and Machine Intelligence*, 33(1), 117–128.
- Johnson, J., Douze, M., & Jégou, H. (2019). Billion-Scale Similarity Search with GPUs. *IEEE Transactions on Big Data*, 7(3), 535–547.
- Babenko, A., & Lempitsky, V. (2014). The Inverted Multi-Index. *IEEE Transactions on Pattern Analysis and Machine Intelligence*, 37(6), 1247–1260.
- Ge, T., He, K., Ke, Q., & Sun, J. (2014). Optimized Product Quantization. *IEEE Transactions on Pattern Analysis and Machine Intelligence*, 36(4), 744–755.
- Douze, M., Guzhva, A., Deng, C., Johnson, J., Szilvasy, G., Mazaré, P.-E., Lomeli, M., Hosseini, L., & Jégou, H. (2024). The Faiss Library. *arXiv preprint arXiv:2401.08281*.
